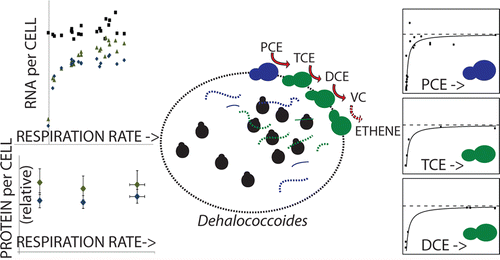
How fast will you eat that chlorinated organic solvent?
Some of the main questions in bioremediation include:
Are the microbes eating (or breathing) that pollutant I want them to?
Are they going to consume it all?
How long is it going to take, and/or can I make it go faster?
It turns out these are more or less central questions in environmental microbiology as well, and there are not great ways to address them! Most techniques in environmental microbiology center around identifying organisms, and unfortunately in the microbial world, that part of their evolutionary history (where microbes got their ribosomes from), may not tell us very much about what they are functionally capable of doing, or if they are actively utilizing that process in that environment. As a graduate student this work was my first exposure to some very real, and long standing challenges in Environmental microbiology. I developed a further appreciation for this working on biomarkers—biological molecules that have the potential to serve as indicators of activity. To do this quantitatively, we had to develop methods of quantifying biomarkers, and incorporate them into activity models, both kinetic and empirical. Specifically, these models looked at how fast an reaction will go in the presence of substrates (food) and amount of biomass (microbes, proteins, etc.). We built models of microbes using DNA (Heavner et al. 2013), and using RNA and Proteins (Rowe et al. 2012 & 2013), all of which were done in mixed microbial community as a model system for a ground water anaerobic community. I’m now utilizing these same techniques and approaches to understanding pollutants and contaminant microbiology impacting ground water in the Ohio river basin. In collaboration with Reza Soltanian and with funding from the Army office of Research, we are investigating not just if there are microbes to remediate contaminants, but what microbes are doing in these systems as they change over time.
Astrobiology, Bioremediation, Electromicrobiology, Environmental Microbiology, Geomicrobiology, and Microbial Ecology.
